Welcome to the TransmiR v3.0
TransmiR is a database for transcription factor (TF)-microRNA (miRNA) regulations, through which one can find regulatory relations between TFs and miRNAs. We updated and presented TransmiR v2.0 in 2018, which included 3,730 TF-miRNA regulation entries from literatures. In this major update to TransmiR v3.0, we integrated increasing regulation data and improved the functionality of databse. Currently, TransmiR v3.0 contains 5,095 literature-curated TF-miRNA regulations, which cover ~946 TFs, ~1,072 miRNAs, 29 organisms and 2,285 publications. Besides, we also provide 6,366,938 TF-miRNA regulations derived from ChIP-seq evidence in 5 species. All TF-miRNA regulations were annotated in detail. If the user have single miRNA, TF or disease at hand, all above transcriptional regulation data can be searched and graphically visualized in "Search" page and "Network" page, respectively. In Network module, users can obtain the disease-specific and sex-dependant TF-miRNA regulation netowrks. Furthermore, we constructed "Enrichment analysis" module to identify the significant TFs that are likely to regulate a miRNA list of interest. Finally, a "Predict" module was implemented, which enables the prediction of TF-miRNA regulations based on binding motif matrices of human TFs. TransmiR v3.0 is freely available for academic usage. Users can also download the datasets for further analysis.
This website has been tested by using Chrome, Microsoft Edge and Firefox browsers. Microsoft IE may not work well.
History:
Aug, 2024, TransmiR v3.0 was released.
May, 2018, TransmiR v2.0 was released.
January 30, 2013, TransmiR v1.2 was released (125 new entries were added into TransmiR).
September, 2009, TransmiR v1.0 was released.
Contact us
Dr. Qinghua Cui
Address: School of Sports Medicine, Wuhan Sports University
Alternative Address: 38 Xueyuan Rd, Department of Biomedical Informatics, Peking University Health Science Center, Beijing 100191, China
Email: cuiqinghua@bjmu.edu.cn
Homepage: http://www.cuilab.cn/
You can search the entries by such keywords:
In the "Search" result, each entry has 10 items.
(1) TF name
(2) miRNA name
(3) TSS
(4) Binding site
(5) Action type
(6) SRAID/PMID
(7) Evidence
(8) Tissue
(9) Species
(10) Details
|
|
|
You can search the entries by such keywords:
In the "Search" result, each entry has 11 items.
(1) TF name
(2) miRNA name
(3) TSS
(4) Regulation start
(5) Regulation end
(6) Score
(7) P-value
(8) Q-value
(9) Evidence
(10) Source
(11) Species
The TransmiR v3.0 data are available as shown below:
- download literature-curated TF-miRNA regulation data here (xls or txt)
- download all evidence level TF-miRNA regulation data based on different species
- download all predicted TF-miRNA regulation data here (tsv.gz)
- download the conserved TF-miRNA regulations (derived from literature) between human and mouse here xls
- The retrieved literature PMID: those used in TransmiR v3.0 and those excluded by TransmiR v3.0
C.elegans (tsv.gz)
D.melanogaster (tsv.gz)
H.sapiens (tsv.gz)
M.musculus (tsv.gz)
R.norvegicus (tsv.gz)
TransmiR v3.0 tutorial (FAQs)
- What are TransmiR, TransmiR v2.0 and TransmiR v3.0?
- How to use "Search" function?
- How to use "Network" function?
- How to use "Enrichment analysis" function?
- How to use "Predict" function?
- How to download datasets?
- What does transcription factor annotation information include?
- What does microRNA annotation information include?
- What is the difference between the evidence level 1 and level 2?
- Which genome assemblies were used in TransmiR v3.0?
Q1. What are TransmiR v2.0 and TransmiR v3.0?
MicroRNAs(miRNAs) are important post-transcriptional regulators of gene expression and play critical roles in various biological processes. It has been reported that aberrant regulation of miRNAs was associated with the development and progression of various diseases, including cancer, cardiovascular disease, etc. But the underlying mechanisms remain elusive, partly because the integrative data of transcription factor (TF)-miRNA regulations is far from sufficient.
TransmiR is a database for transcription factor (TF)-microRNA (miRNA) regulations, through which one can find regulatory relations between TFs and miRNAs. TransmiR was originally constructed on September 2009 and updated on January 2013. In this major update to TransmiR v2.0, we manually curated 2,852 TF-miRNA regulations from 1,045 publications during 2013-2017 and included ChIP-seq-derived TF-miRNA regulation data. Currently, TransmiR v2.0 contains 3,730 literature-curated TF-miRNA regulations, which cover ~623 TFs, ~785 miRNAs, 19 organisms and 1,349 publications. Besides, we also provide 1,785,998 TF-miRNA regulations derived from ChIP-seq evidence in 5 species.
TransmiR v3.0 database is presented with more comprehensive miRNA transcription regulation information, which contains 5,110 transcription factor (TF) -miRNA regulations curated from 2,291 literatures and more than 6 million TF-miRNA regulations derived from ChIP-seq data. Currently, TransmiR v3.0 covers 3,260 TFs, 4,256 miRNAs, and 514,423 TF-miRNA regulation pairs among 29 organisms. Additionally, motif scanning of TF loci on promoter sequences of miRNAs from multiple species is employed to predict the TF-miRNA regulations, generating 284,527 predicted TF-miRNA regulations. Besides the significant growth of data volume, we also improve the annotation for TFs and miRNAs by introducing TF family, TFBS motif, and expression profiles for several species. Moreover, functionality of the TransmiR v3.0 online database is enhanced, including flexible query supporting batch searching, and more extensive disease-specific, as well as newly sex-specific TF-miRNA regulation networks in ‘Network’ modules.TransmiR v3.0 is freely available for academic usage. Users can also download the datasets for further analysis.
Q2. How to use "Search" function?
To search data in the database by a miRNA or TF name, please click the "Search" tab in the menu (1 in Figure 1). TransmiR v3.0 provides functions to "Search" with two modes (i.e single and multiple keywords mode, respectively) of transcription factors and microRNAs. Specify the species, input type (TF or miRNA) and searching mode (single keyword or multiple keywords) (2 in Figure 1), then you can input your search keywords (i.e "hsa-mir-200a") into corresponding blanks (3 in Figure 1) and submit the query (4 in Figure 1). In single keyword mode, the tips for users' input will be poped out (Figure 2), thereby users can select the truly interested one. And the multiple keyword mode allows users to input multiple TFs or miRNAs split by ‘;’ (Figure 3).
Each "search" result will return entries including the following nine items.
(1) TF name - The link to detailed information for the transcription factor;
(2) miRNA name - The link to detailed information for the miRNA;
(3) TSS - microRNA gene transcription start site;
(4) TF binding site - Genomic coordinates of TF binding sites derived from ChIP-seq data;
(5) Action type - Indicating the regulatory type: Activation or Repression (when the action type is unclear, shown as "Regulation"). And for each entry, "(feedback)" was added if the TF turns out to be the target gene of the miRNA it regulates. Such feedback miRNA-TF regulations are derived from the experimentally validated miRNA target genes in TarBase or miRTarBase;
(6) SRAID/PMID - The PMID links to the reference in PubMed for literature-curated regulations, or the SRA IDs for ChIP-seq derived regulations;
(7) Evidence - The confidence level of this TF-miRNA regulation, including level 1, level 2 and literature. All literature-curated TF-miRNA regulations were annotated as "literature". Based on reliability of the promoter region annotation used, we classified the TF-miRNA regulations derived from ChIP-seq into level 1 and level 2 (level 2 promoter is supported by high-throughput experimental data, see more in Q.9 in FAQs.);
(8) Tissue - The tissue from which the ChIP-seq data was derived;
(9) Species - The species of the TF-miRNA regulation.
You can start a new search by clicking the "Reset all" button (5 in Figure 1). And you can also download the search result by clicking "here" (6 in Figure 1).
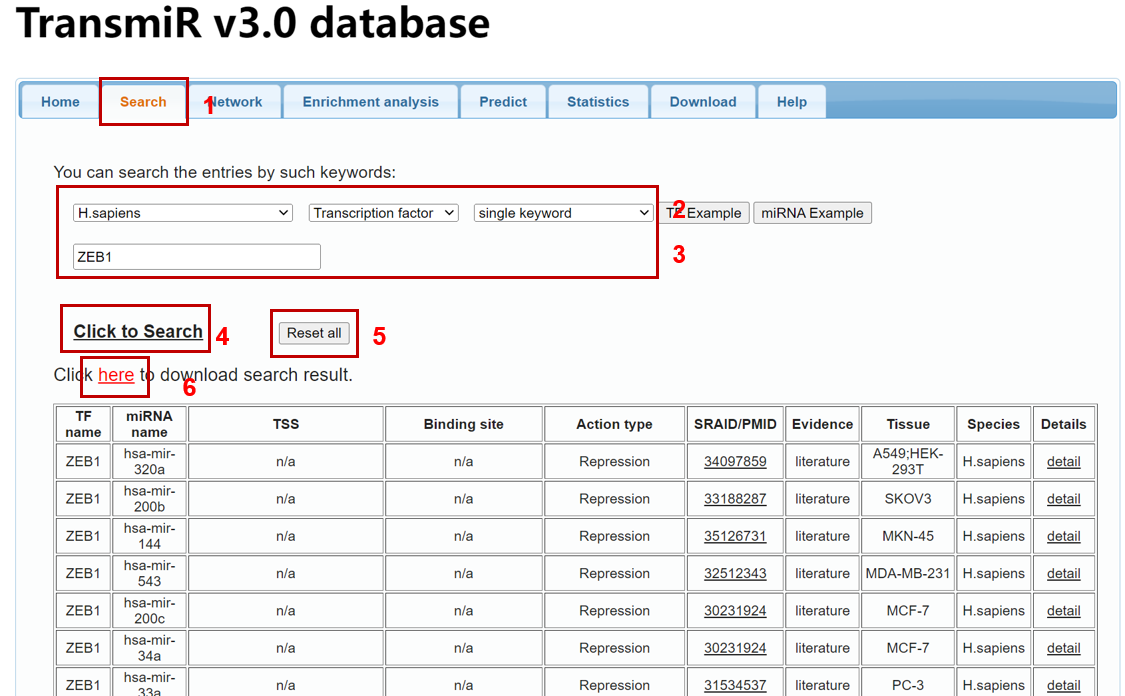
Figure 1. A demo for searching TF-miRNA regulations in the database
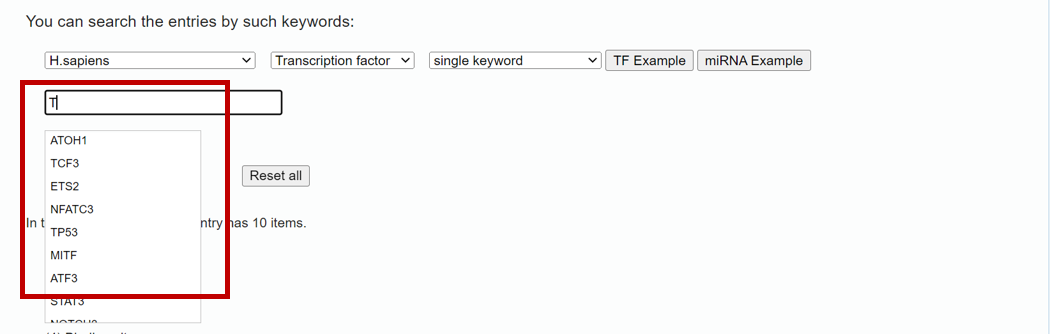
Figure 2. A demo for single keyword search mode
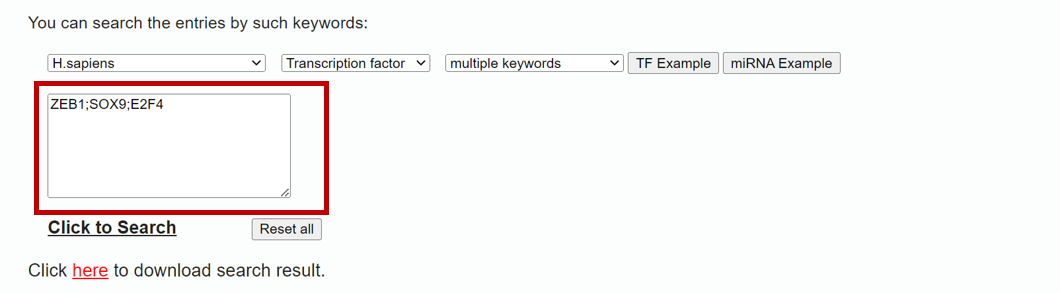
Figure 3. A demo for single keywords search mode
Q3. How to use "Network" function?
To visualize TF-miRNA regulation data in the form of a network diagram, please click the "Network" tab in the menu (1 in Figure 4). To browse the TF-miRNA regulatory network related to one miRNA, please click "miRNA" to expand the tree architecture, then choose the species and select the specific miRNA you are interested in. The network related to specific TF or disease can be browsed by the similar way. For example, if you want to view the TF-miRNA regulatory network related to "Diabetes Mellitus", please click the "disease" (2 in Figure 4), and then select "Diabetes Mellitus" (3 in Figure 4). In addition, users have the option to select a confidence level for filtering TF-miRNA regulations (4 in Figure 4), including literature-derived TF-miRNA regulations, level 2 (promoter regions originating from high-throughput data), level 1, and all regulations. The default disease network is composed of literature-derived TF-miRNA regulation relationships. The corresponding network diagram will be shown on the right panel (5 in Figure 4). The legend of the network is available at the bottom-right corner of the panel (6 in Figure 4), where the meaning of each element in the network is explained. Users can download the disease regulation result (7 in Figure 4). Each TF-miRNA regulatory network is constituted by TFs and miRNAs as its nodes, while TF-miRNA regulations and miRNA-TF regulations as its edges. Within the network view, you can drag a node with the mouse and use "mouse wheel rolling" operation to zoom in/out. Moreover, the linked nodes are highlighted when a node is clicked (8 in Figure 4). Firefox, Safari, IE9, and Chrome can all display the TF-miRNA regulatory network properly.
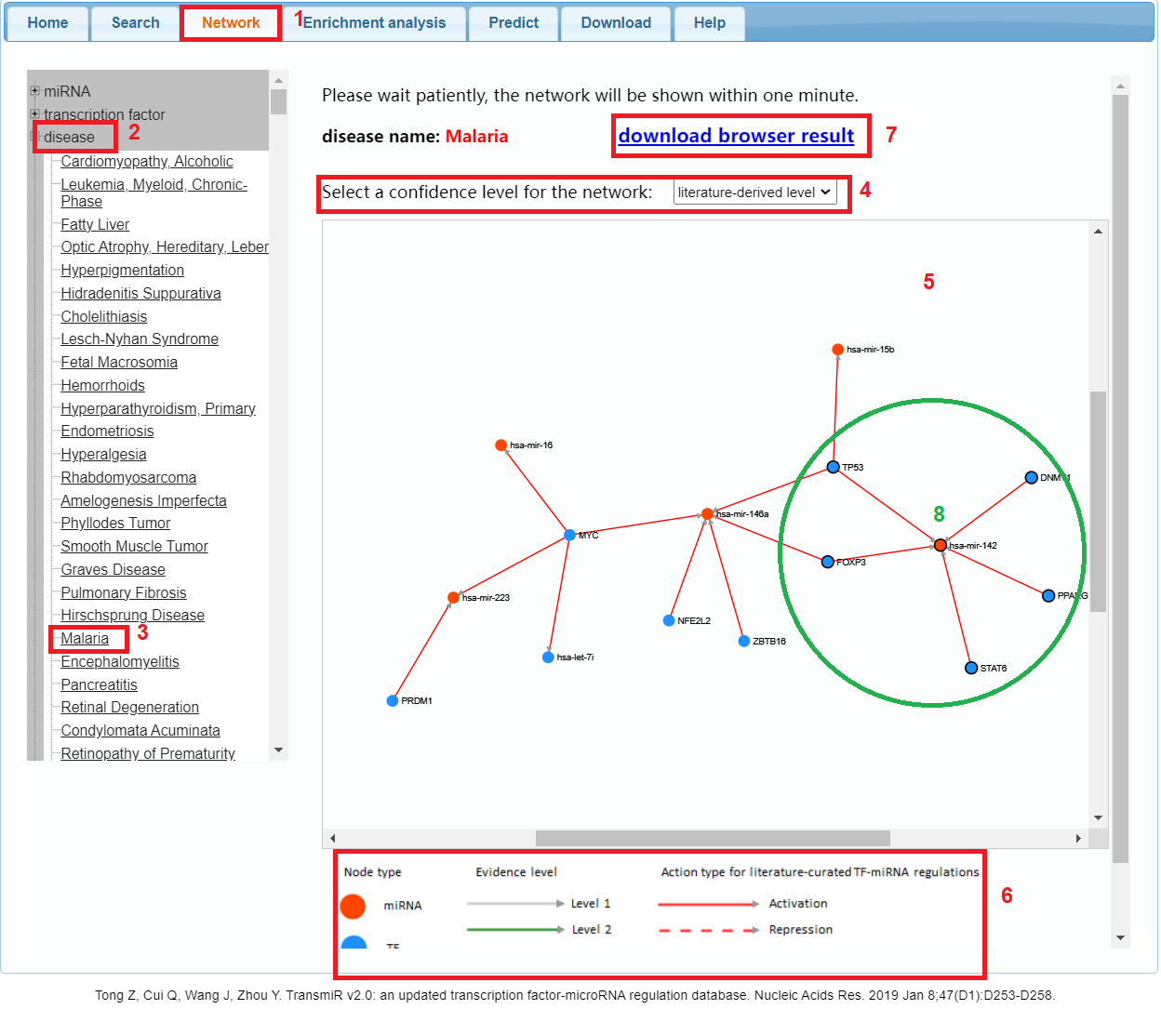
Figure 4. A demo for browsing TF-miRNA regulatory network in the database
Q4. How to use "Enrichment analysis" function?
If you have a list of miRNAs and want to find the overrepresented TFs regulate them, please click the "Enrichment analysis" in the menu (1 in Figure 5). The enrichment (overrepresentation) analysis of TFs can be performed as described below. Firstly, input or paste a list of miRNAs into the text box (2 in Figure 5), you can also click "Example" button (3 in Figure 5) to load an example miRNA list. Secondly, set the size of miRNA category (4 in Figure 5). This choice limits the size of miRNA category to a certain range. Thirdly, choose the lowest confidence level (5 in Figure 5). Level 2 data is limited to the miRNA promoters supported by experimental data, see more in Q.9 in FAQs. Therefore, if you prefer better stringency but less coverage of the analysis results, you should choose "Set level 2" option. Finally, click the "run" button (6 in Figure 5) to submit the query. You can browse the result of enrichment analysis in a new tabular page, where the miRNA category (i.e. regulating TF) and the associated statistics (P-value, Bonferroni, FDR, etc.) from Hypergeometric distribution analysis are shown.

Figure 5. A demo for analyzing the overrepresented regulating transcription factors for a miRNA list of interest
Q5. How to use "Predict" function?
To query the predicted TF-miRNA regulations based on binding motif matrices of human TFs, please click the "Predict" tab in the menu (1 in Figure 6). "Predict" module provides functions to query the prediction results by full/partial names (i.e exact and fuzzy modes, respectively) of transcription factors and microRNAs. Specify input type (TF or miRNA) and searching mode (exact or fuzzy) (2 in Figure 6), then you can input your search keywords (i.e "ZEB1") into corresponding blanks (3 in Figure 6) and submit the query (4 in Figure 6).
Each "search" result will return entries including the following eight items.
(1) TF name; (2) miRNA name; (3) TSS, i.e. transcriptional start site of miRNA;
(4) Start - The regulation start postion in corresponding miRNA promoter region.
(5) End - The regulation end postion in corresponding miRNA promoter region.
(6) Evidence - The confidence level of this TF-miRNA regulation, including level 1 and level 2. Based on reliability of the promoter region annotation used, we classified the TF-miRNA regulations derived from ChIP-seq into level 1 and level 2 (level 2 promoter is supported by high-throughput experimental data, see more in Q.9 in FAQs.);
(7) Score - which is computed by summing the appropriate entries from each column of the position-dependent scoring matrix that represents the motif;
(8) P-value - the probability of a random sequence of the same length as the motif matching that position of the sequence with as good or better a score;
(9) Q-value - the false discovery rate if the occurrence is accepted as significant.;
(10) Source - The soure database of TF, including JASPAR and HOCOMOCO;
(11) Species - which includs human, mouse and rat;
You can start a new search by clicking the "Reset all" button (5 in Figure 6). And you can also download the search result by clicking "here" (6 in Figure 6).
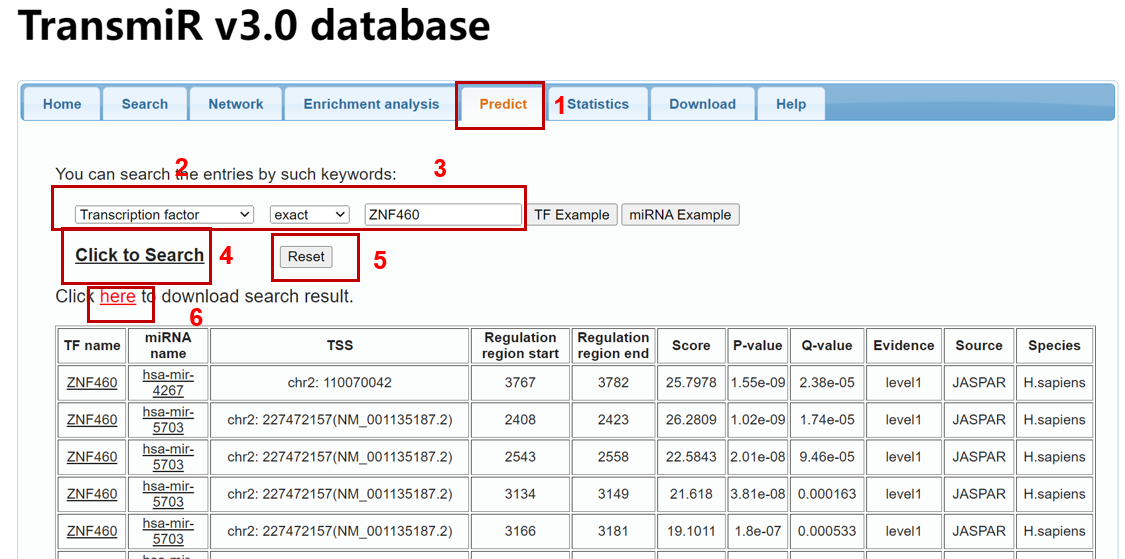
Figure 6. A demo for searching predicted TF-miRNA regulations in the database
Q6. How to download datasets?
To download datasets in the database, please click the "Download" tab in the menu. TransmiR v3.0 provides two formats of downloadable files, i.e. Excel .xls and textual .tsv formats, respectively. The users can download both literature-curated TF-miRNA regulation data, all TF-miRNA regulation data (including the data derived from ChIP-seq experiments), and all predicted TF-miRNA regulation data.
Q7. What does transcription factor annotation information include?
Each transcription factor annotation entry contains the following five items: Gene official symbol, Entrez gene ID, Ensembl gene ID, TF family (from AnimalTFDB and PlantTFDB), gene associated diseases (from DisGeNet database), cancer prognostic association data (from The Human Protein Atlas database), and TFBS motif (from JASPAR 2024 and HOCOMOCO v12). And TF gene expression profiles in normal tissues (from BGee database) and cancer tissues (from TCGA database).
Q8. What does miRNA annotation information include?
Each miRNA annotation entry contains the following five items: miRNA name, miRBase ID, genome context of this miRNA gene, miRNA associated diseases from HMDD v4.0 database and a link to miRBase database. And miRNA gene expression profiles in normal tissues (from BGee database) and cancer tissues (from TCGA database).
Q9. What is the difference between the evidence level 1 and level 2?
Based on reliability of the transcriptional regulatory region (promoter region) annotation used, we classified the TF-miRNA regulations derived from ChIP-seq into level 1 and level 2. For level 1, we choose 5'-end of the pre-miRNA or that of the first member in the miRNA cluster as the transcription start site. Next, a window from the 5kb upstream to the 1kb downstream of the miRNA TSS was identified as the putative transcriptional regulatory region. Apparently, this definition could cover most of the miRNAs, but suffers from substantial inaccuracy. Therefore for level 2, the miRNA TSS was obtained from FANTOM v6. And the 300bp upstream and 100bp downstream of each miRNA TSS was identified as the putative transcriptional regulatory region. The level 2 TF-miRNA regulations are much more stringent than level 1 TF-miRNA regulations, but cover less miRNAs.
Q10. Which genome assemblies were used in TransmiR v3.0?
The genome assembly versions of all species are consistent with miRBase v22. The specific genome assembly versions are as follows.
- hg38 (H.sapiens)
- mm10 (M.musculus)
- rn6 (R.norvegicus)
- ce11 (C.elegans)
- dm6 (D.melanogaster)
If there is any comment or suggestion, please contact us:
Dr. Chunmei Cui (ccm328@bjmu.edu.cn)
Dr. Qinghua Cui (cuiqinghua@bjmu.edu.cn)